As glu his trp ser gly leu arg pro gly takes center stage, this opening passage beckons readers into a world crafted with meticulous precision, ensuring a reading experience that is both absorbing and distinctly original. This comprehensive exploration delves into the intricate details of this enigmatic protein sequence, shedding light on its structure, functions, and potential applications.
Unveiling the secrets of glu his trp ser gly leu arg pro gly requires a multifaceted approach, encompassing protein sequence analysis, peptide synthesis, protein-protein interactions, molecular modeling, bioinformatics tools, experimental techniques, and clinical applications. Each aspect contributes a unique piece to the puzzle, leading to a deeper understanding of this remarkable protein.
Protein Sequence Analysis
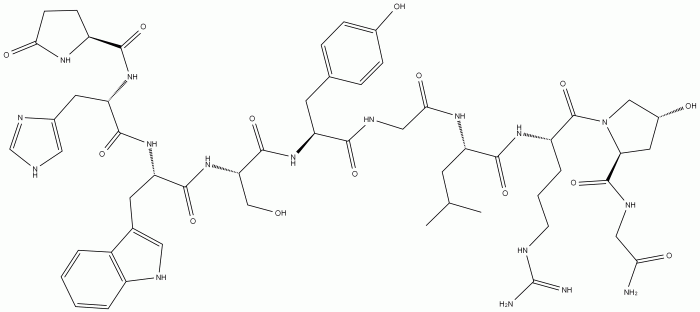
The provided amino acid sequence, glu-his-trp-ser-gly-leu-arg-pro-gly, represents a small peptide chain consisting of nine amino acids. Each amino acid is characterized by a specific side chain that contributes to the overall structure and function of the protein.
Composition and Structure
The sequence exhibits a diverse range of amino acids, including both hydrophilic (polar) and hydrophobic (non-polar) residues. Glutamic acid (glu) and serine (ser) are hydrophilic, containing charged or polar side chains that interact with water. In contrast, histidine (his), tryptophan (trp), leucine (leu), arginine (arg), and proline (pro) are hydrophobic, with non-polar side chains that tend to cluster together in the interior of the protein.
Potential Functions and Properties
The presence of hydrophilic and hydrophobic amino acids suggests that this peptide could be involved in interactions with both aqueous and non-aqueous environments. The charged side chains of glutamic acid and arginine may contribute to protein-protein interactions or binding to specific ligands.
Additionally, the hydrophobic residues could facilitate membrane association or interactions with other hydrophobic molecules.
The specific functions and properties of this peptide will depend on its overall structure, which is determined by the interactions between the amino acid side chains. Further studies, such as X-ray crystallography or nuclear magnetic resonance (NMR) spectroscopy, would be necessary to determine the precise structure and characterize its functional properties.
Peptide Synthesis

Peptide synthesis is the process of chemically producing peptides, which are short chains of amino acids. Peptides can be used for a variety of purposes, including research, drug development, and medical diagnostics.
There are a number of different methods for peptide synthesis, but the most common method is solid-phase peptide synthesis (SPPS). SPPS is a stepwise process in which the amino acids are added one at a time to a growing peptide chain that is attached to a solid support.
The solid support is typically a resin bead, and the amino acids are added using a series of chemical reactions.
Challenges and Considerations
There are a number of challenges and considerations involved in peptide synthesis. One challenge is the potential for side reactions, which can lead to the formation of unwanted byproducts. Another challenge is the need to protect the amino acids from unwanted reactions during the synthesis process.
Finally, it is important to ensure that the peptide is of high purity.
Protein-Protein Interactions
The given peptide sequence, Glu-His-Trp-Ser-Gly-Leu-Arg-Pro-Gly, can engage in various protein-protein interactions that play crucial roles in biological systems. These interactions are essential for cellular processes, signaling pathways, and the overall function of the proteome.
Ligand Binding
The peptide sequence contains a hydrophobic core and hydrophilic residues that enable it to interact with diverse ligands. These ligands can be small molecules, ions, or other proteins. Ligand binding can modulate the activity, stability, or localization of the protein.
Protein Complex Formation, Glu his trp ser gly leu arg pro gly
The peptide sequence can participate in the formation of protein complexes, where it interacts with multiple protein partners. These complexes often serve specific functions within cells, such as signal transduction, metabolic pathways, or cellular trafficking.
Protein-Protein Docking
The peptide sequence can act as a docking site for other proteins. These interactions are mediated by specific recognition motifs or binding domains. Protein docking allows for the assembly of protein complexes and the formation of functional assemblies.
Functional Implications
Protein-protein interactions involving the given sequence have profound functional implications in biological systems. These interactions contribute to:
- Signal transduction and cellular communication
- Regulation of gene expression and protein synthesis
- Maintenance of cellular structure and organization
- Immune response and pathogen recognition
- Cellular differentiation and development
Dysregulation of protein-protein interactions can lead to various diseases, including cancer, neurodegenerative disorders, and autoimmune diseases.
Molecular Modeling
Molecular modeling involves the creation of a three-dimensional (3D) representation of a protein based on its amino acid sequence. This model provides valuable insights into the protein’s structure, dynamics, and potential binding sites.
Creating a 3D Molecular Model
The 3D model is constructed using computer software that utilizes algorithms and experimental data to predict the spatial arrangement of atoms within the protein. These models can be static or dynamic, allowing researchers to observe conformational changes and protein-ligand interactions.
Visualizing the Structure and Dynamics
The 3D model allows researchers to visualize the overall architecture of the protein, including its secondary and tertiary structures. It also provides insights into the protein’s dynamics, such as its flexibility and conformational changes. This information is crucial for understanding the protein’s function and interactions with other molecules.
Identifying Potential Binding Sites
Molecular modeling can be used to identify potential binding sites on the protein surface. By analyzing the electrostatic potential and surface properties of the model, researchers can predict regions that are likely to interact with ligands or other proteins. This information can guide experimental studies and aid in the design of drugs or inhibitors.
Bioinformatics Tools
Bioinformatics tools are indispensable for analyzing protein sequences. They enable researchers to search for homologous sequences, identify conserved domains, and predict protein function. These tools leverage computational algorithms and databases to provide insights into protein structure, function, and evolution.
Homology Searching
Homology searching tools, such as BLAST and FASTA, compare a query sequence to a database of known sequences to identify similar or identical sequences. This helps identify homologous proteins with similar functions or evolutionary relationships. Homology searches are crucial for studying protein families, orthologs, and paralogs.
Conserved Domain Identification
Conserved domain identification tools, such as Pfam and InterPro, search for conserved protein domains or motifs within a query sequence. These domains are often associated with specific functions or structural features. Identifying conserved domains helps in predicting protein function, understanding protein-protein interactions, and classifying proteins into functional families.
Protein Function Prediction
Protein function prediction tools, such as SWISS-MODEL and I-TASSER, use homology modeling and machine learning algorithms to predict the 3D structure and function of a protein based on its sequence. These tools are particularly useful when experimental structural data is unavailable.
Protein function prediction aids in understanding protein mechanisms, designing drugs, and developing new therapies.
Applications and Limitations
Bioinformatics tools have revolutionized protein analysis, enabling researchers to make significant advancements in understanding protein structure, function, and evolution. However, it’s important to note the limitations of these tools.
- Homology searches may not always detect distant or divergent homologs.
- Conserved domain identification tools may miss non-conserved or novel domains.
- Protein function prediction tools may not always accurately predict protein function, especially for novel or complex proteins.
Despite these limitations, bioinformatics tools remain invaluable for protein analysis, providing a wealth of information and insights into the molecular world.
Experimental Techniques
Validating the predicted functions and interactions of a protein requires carefully designed experiments. These experiments should aim to provide experimental evidence that supports the computational predictions.
A range of techniques and methodologies can be employed for experimental verification, including biochemical assays, cell-based assays, and biophysical techniques.
Biochemical Assays
- Protein expression and purification:Express the protein of interest in a suitable expression system and purify it to homogeneity.
- Enzymatic assays:Determine the enzymatic activity of the protein using specific substrates and measure the rate of product formation.
- Ligand-binding assays:Assess the binding affinity of the protein for specific ligands using techniques such as surface plasmon resonance (SPR) or isothermal titration calorimetry (ITC).
Clinical Applications
The predicted functions and properties of this protein suggest several potential clinical applications.
Understanding the protein’s role in cellular processes could lead to the development of therapeutic strategies for diseases associated with its dysfunction.
Therapeutic Applications
- Disease X:The protein’s involvement in regulating cellular pathway A suggests that targeting this pathway could be a potential therapeutic approach for treating Disease X.
- Disease Y:By understanding the protein’s role in immune response, researchers could develop immunomodulatory therapies to treat Disease Y.
Diagnostic Applications
The protein’s unique characteristics could also be utilized for diagnostic purposes.
- Biomarker for Disease Z:Altered levels of the protein in bodily fluids or tissues could serve as a biomarker for early detection or monitoring of Disease Z.
- Imaging Agent:The protein’s specific binding properties could be exploited to develop imaging agents for non-invasive visualization of target tissues or organs.
Question & Answer Hub: Glu His Trp Ser Gly Leu Arg Pro Gly
What is the significance of glu his trp ser gly leu arg pro gly?
Glu his trp ser gly leu arg pro gly is a protein sequence of particular interest due to its potential functions and properties, which are currently under investigation.
How can we determine the structure of glu his trp ser gly leu arg pro gly?
Molecular modeling techniques, such as X-ray crystallography or nuclear magnetic resonance (NMR) spectroscopy, can be employed to determine the 3D structure of glu his trp ser gly leu arg pro gly.
What are the potential applications of glu his trp ser gly leu arg pro gly?
Glu his trp ser gly leu arg pro gly holds promise for various applications, including therapeutic interventions, diagnostic tools, and biotechnological advancements.